Dask Read Parquet Files into DataFrames with read_parquet#
This blog post explains how to read Parquet files into Dask DataFrames. Parquet is a columnar, binary file format that has multiple advantages when compared to a row-based file format like CSV. Luckily Dask makes it easy to read Parquet files into Dask DataFrames with read_parquet.
It’s important to properly read Parquet files to take advantage of performance optimizations. Disk I/O can be a major bottleneck for distributed compute workflows on large datasets. Reading Parquet files properly allows you to send less data to the computation cluster, so your analysis can run faster.
Let’s look at some examples on small datasets to better understand the options when reading Parquet files. Then we’ll look at examples on larger datasets with thousands of Parquet files that are processed on a cluster in the cloud.
Dask read_parquet: basic usage#
Let’s create a small DataFrame and write it out as Parquet files. This will give us some files to try out read_parquet. Start by creating the DataFrame.
import dask.dataframe as dd
import pandas as pd
df = pd.DataFrame(
{"nums": [1, 2, 3, 4, 5, 6], "letters": ["a", "b", "c", "d", "e", "f"]}
)
ddf = dd.from_pandas(df, npartitions=2)
Now write the DataFrame to Parquet files with the pyarrow engine. The installation instructions for pyarrow are in the Conda environment for reading Parquet files section that follows.
ddf.to_parquet("data/something", engine="pyarrow")
Here are the files that are output to disk.
data/something/
_common_metadata
_metadata
part.0.parquet
part.1.parquet
You can read the files into a Dask DataFrame with read_parquet.
ddf = dd.read_parquet("data/something", engine="pyarrow")
Check the contents of the DataFrame to make sure all the Parquet data was properly read.
ddf.compute()

Dask read Parquet supports two Parquet engines, but most users can simply use pyarrow, as we’ve done in the previous example, without digging deep into this option.
Dask read_parquet: pyarrow vs fastparquet engines#
You can read and write Parquet files to Dask DataFrames with the fastparquet and pyarrow engines. Both engines work fine most of the time. The subtle differences between the two engines doesn’t matter for the vast majority of use cases.
It’s generally best to avoid mixing and matching the Parquet engines. For example, you usually won’t want to write Parquet files with pyarrow and then try to read them with fastparquet.
This blog post will only use the pyarrow engine and won’t dive into the subtle differences between pyarrow and fastparquet. You can typically just use pyarrow and not think about the minor difference between the engines.
Dask read_parquet: lots of files in the cloud#
Our previous example showed how to read two Parquet files on localhost, but you’ll often want to read thousands of Parquet files that are stored in a cloud based file system like Amazon S3.
Here’s how to read a 662 million row Parquet dataset into a Dask DataFrame with a 5 node computational cluster.
import coiled
import dask
import dask.dataframe as dd
cluster = coiled.Cluster(name="read-parquet-demo", n_workers=5)
client = dask.distributed.Client(cluster)
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
)
Take a look at the first 5 rows of this DataFrame to get a feel for the data.
ddf.head()
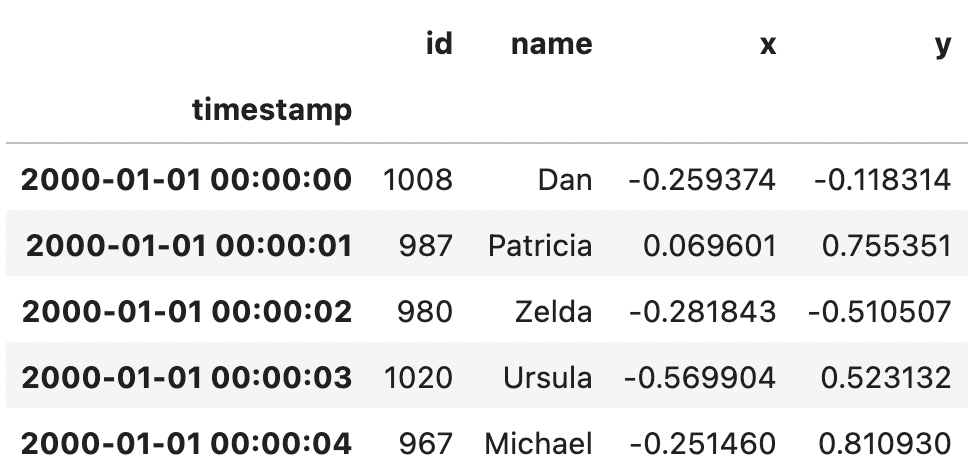
This dataset contains a timestamp index and four columns of data.
Let’s run a query to compute the number of unique values in the id column.
ddf["id"].nunique().compute()
This query takes 59 seconds to execute.
Notice that this query only requires the data in the id column. However, we transferred the data for all columns of the Parquet file to run this query. Spending time to transfer data that’s not used from the filesystem to the cluster is obviously inefficient.
Let’s see how Parquet allows you to only read the columns you need to speed up query times.
Dask read_parquet: column selection#
Parquet is a columnar file format which allows you to selectively read certain columns when reading files. You can’t cherry pick certain columns when reading from row-based file formats like CSV. Parquet’s columnar nature is a major advantage.
Let’s refactor the query from the previous section to only read the id column to the cluster by setting the columns argument.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
columns=["id"],
)
Now let’s run the same query as before.
ddf["id"].nunique().compute()
This query only takes 43 seconds to execute, which is 27% faster. This performance enhancement can be much larger for different datasets / queries.
Cherry picking individual columns from files is often referred to as column pruning. The more columns you can skip, the more column pruning will help speed up your query.
Definitely make sure to leverage column pruning when you’re querying Parquet files with Dask.
Dask read_parquet: row group filters#
Parquet files store data in row groups. Each row group contains metadata, including the min/max value for each column in the row group. For certain filtering queries, you can skip over entire row groups just based on the row group metadata.
For example, suppose columnA in row_group_3 has a min value of 2 and a max value of 34. If you’re looking for all rows with a columnA value greater than 95, then you know row_group_3 won’t contain any data that’s relevant for your query. You can skip over the row group entirely for that query.
Let’s run a query without any row group filters and then run the same query with row group filters to see the performance book predicate pushdown filtering can provide.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
)
len(ddf[ddf.id > 1170])
This query takes 77 seconds to execute.
Let’s run the same query with row group filtering.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
filters=[[("id", ">", 1170)]],
)
len(ddf[ddf.id > 1170])
This query runs in 4.5 seconds and is significantly faster.
Row group filtering is also known as predicate pushdown filtering and can be applied in Dask read Parquet by setting the filters argument when invoking read_parquet.
Predicate pushdown filters can provide massive performance gains or none at all. It depends on how many row groups Dask will be able to skip for the specific query. The more row groups you can skip with the row group filters, the less data you’ll need to read to the cluster, and the faster your analysis will execute.
Dask read_parquet: ignore metadata file#
When you write Parquet files with Dask, it’ll output a _metadata file by default. The _metadata file contains the Parquet file footer information for all files in the filesystem, so Dask doesn’t need to individually read the file footer for every file in the Parquet dataset every time the Parquet lake is read.
The _metadata file is a nice performance optimization for smaller datasets, but it has downsides.
_metadata is a single file, so it’s not scalable for huge datasets. For large data lakes, even the metadata can be “big data”, with the same scaling issues of “regular data”.
You can have Dask read Parquet ignore the metadata file by setting ignore_metadata_file=True.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
ignore_metadata_file=True,
)
Dask will gather and process the metadata for each Parquet file in the lake when it’s instructed to ignore the _metadata file.
Dask read_parquet: index#
You may be surprised to see that Dask can intelligently infer the index when reading Parquet files. Dask is able to confirm the index from the Pandas parquet file metadata. You can manually specify the index as well.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
index="timestamp",
)
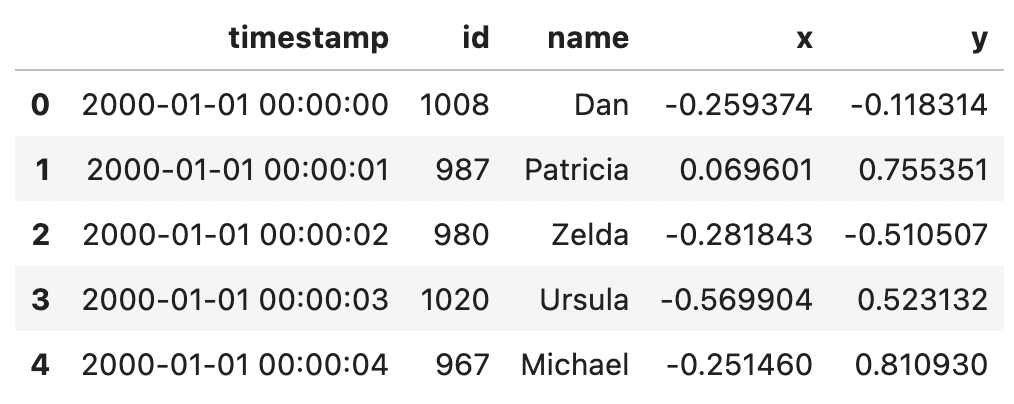
You can also read in all data as regular columns without specifying an index.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
index=False,
)
ddf.head()
Dask read_parquet: categories argument#
You can read in a column as a category column by setting the categories option.
ddf = dd.read_parquet(
"s3://coiled-datasets/timeseries/20-years/parquet",
engine="pyarrow",
storage_options={"anon": True, "use_ssl": True},
categories=["id"],
)
Check the dtypes to make sure this was read in as a category.
ddf.dtypes
id category
name object
x float64
y float64
dtype: object
Conda environment for reading Parquet#
Here’s an abbreviated Conda YAML file for creating an environment with the pyarrow and fastparquet dependencies:
name: standard-coiled
channels:
- conda-forge
- defaults
dependencies:
- python=3.9
- pandas
- dask
- pyarrow
- fastparquet
You don’t need to include both pyarrow and fastparquet in your environment. Just add the Parquet engine you’ll be using.
You can use this environment when running Dask to keep your life simple.
Additional resources#
Here are additional resources if you’d like to learn more:
How to speed up a Pandas query 10x with 6 Dask tricks, many of which involve the Parquet tactics outlined in this post
Conclusion#
This blog post showed you how to properly read Parquet files with Dask.
There are a lot of options and they can impact the runtime of your analysis significantly, so knowing how to read Parquet files is quite important.
Column pruning is of particular importance. It’s easy to apply column pruning and it often yields a significant performance gain.